The potential use of DNA in data storage applications has been in the HEXUS news several times in recent years. One of the big players in the field is Microsoft, which has invested quite a bit of expertise and cash into DNA data storage technology, working alongside the likes of the University of Washington and DNA synthesis company Twist Bioscience.

People and 'things' are generating vast amounts of data every minute, presenting problems for tech companies, especially those with video sharing and cloud features. However, back in 2016 Microsoft and UW researchers set a DNA data storage milestone which they said meant that "all the publicly accessible data on the Internet," could be fitted in a storage medium the size of a shoebox. That sounds fantastic but there are some snags in using DNA storage that needed to be overcome to see it reach its potential.
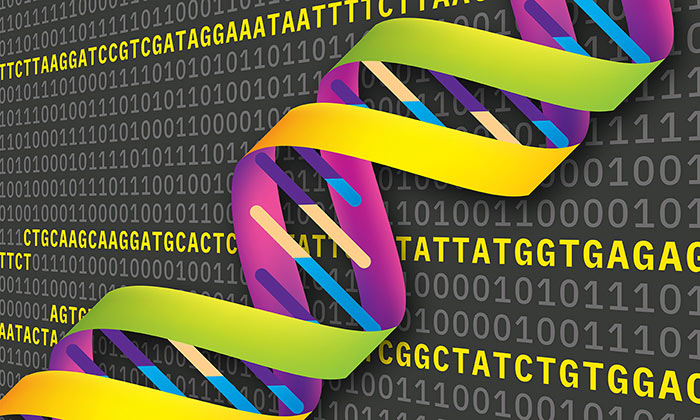
One big problem with DNA storage previously was that there is a naturally a high rate of errors in encoding the information. Thus, to make sure the information was squirreled away safely, DNA encoders would need to repeat the code for any data 10x or even 15x, which is quite inefficient. Now Scientists at the University of Texas at Austin have come up with a way to store information in strands of DNA with novel error correction code.
An algorithm, developed by the research team lead by Ilya J. Finkelstein, tackles the unique characteristics of DNA storage and its 10 per cent error rate. The DNA 'alphabet' isn't binary, it has four bases; A, C, G and T (adenine, cytosine, guanine, and thymine), which facilitates far denser storage but makes error correcting algorithm development much harder. The team built the information recordings on DNA like a lattice, so piece of information reinforces other pieces of information.
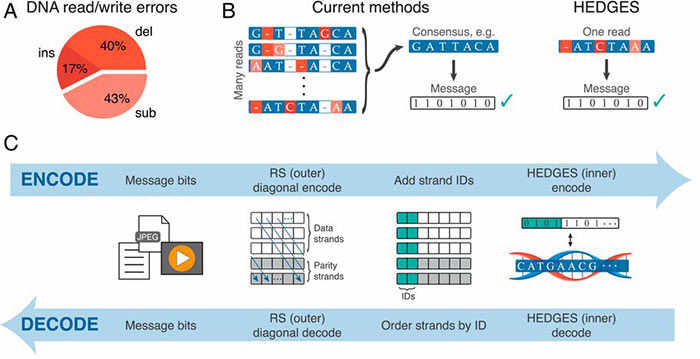
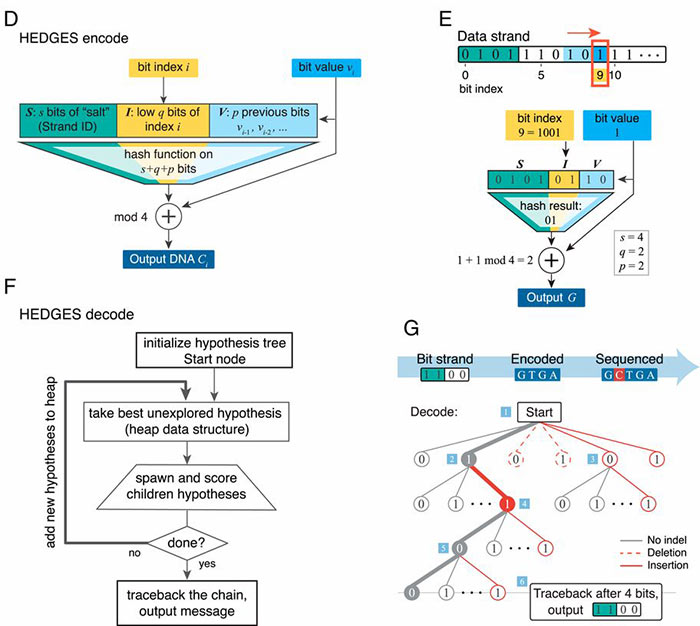
UT researchers have developed HEDGES (Hash Encoded, Decoded by Greedy Exhaustive Search) error-correcting code that repairs all three basic types of DNA errors: insertions, deletions, and substitutions. Moreover, HEDGES converts unresolved or compound errors into substitutions and can avoid excess repeats, or too high or too low windowed guanine–cytosine (GC) content.
To test HEDGES the UT team stored the entirety of The Wizard of Oz, translated into Esperanto, "with more accuracy than prior DNA storage methods ever could have," reports Popular Mechanics. The stored data was subjected to high temperatures and extreme humidity, which damaged the DNA, but "all the information was still decoded successfully," says the UT news blog.
One must remember the potential of DNA - "A teaspoon of DNA contains so much data it would require about 10 Walmart Supercentre-sized data centres to store using current technology," John Hawkins, another co-author of the new paper, told the Popular Mechanics.
So, how long will it be until DNA storage starts to be utilised by scientists or commercial organisations? Finkelstein reckons "niche applications are probably close to being on the horizon, but I don’t think it's going to be a mass market product for a decade or more."













